|
These mutations can also impact the degree to which the protein is glycosylated by creating new or removing existing locations of the glycans (glycosites) on the surface antigens 15, 16. Over time, the protein sequences in viral antigens undergo mutations (antigenic drift), which can alter the species specificity of the virus 12, modulate its infectivity 13, and alter the antigenicity of the surface proteins 14. Additionally, because the virus hijacks the host cellular machinery for replication and subsequent glycosylation, the viral glycan shield may be composed of familiar host glycans thereby suppressing an anti-carbohydrate immune response 9.įortunately, the innate immune system has evolved a range of strategies for responding to glycosylated pathogens 10, but antigen glycosylation nevertheless complicates the development of vaccines 11. The glycosylation of these surface antigens helps the pathogen evade recognition by the host immune system by cloaking the protein surface from detection by the humoral and cellular components of the innate immune system 3, 4, 5, and by altering the ability of the host to raise an effective adaptive immune response 6, 7 or even by enhancing infectivity 8. Viral envelope proteins are often modified by the attachment of complex glycans that can account for up to half of the molecular weight of these glycoproteins, as in HIV gp120 2. Here we examine the structure of the SARS-CoV-2 envelope spike (S) protein that mediates host cell infection, with a specific focus on the extent to which glycosylation masks this virus antigen from the host immune response. As of July 2020, there is still no vaccine or approved therapeutic to treat this disease. The current COVID-19 pandemic has led to over 11 million confirmed infections globally with a fatality rate of approximately 4.5% 1 since the first reports of a severe acute respiratory syndrome (SARS) infection by a novel coronavirus (SARS-CoV-2) at the end of 2019. Despite the relatively modest contribution of the glycans to the total molecular weight of the S trimer (17% for the HEK293 glycoform) they shield approximately 40% of the protein surface. The 3D structures show that the protein surface is extensively shielded from antibody recognition by glycans, with the notable exception of the ACE2 receptor binding domain, and also that the degree of shielding is largely insensitive to the specific glycoform. Lastly, we have identified peptides in the S glycoprotein that are likely to be presented in human leukocyte antigen (HLA) complexes, and discuss the role of S protein glycosylation in potentially modulating the innate and adaptive immune response to the SARS-CoV-2 virus or to a related vaccine. 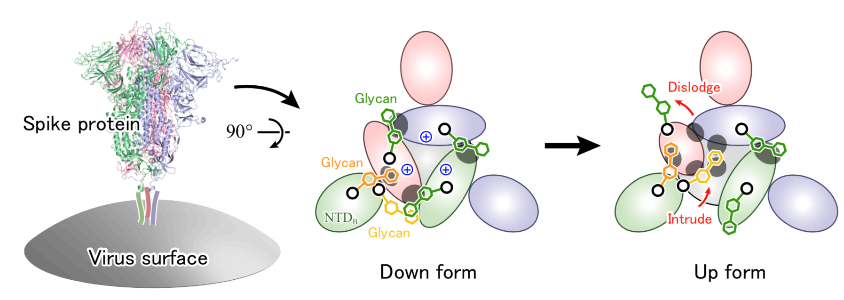 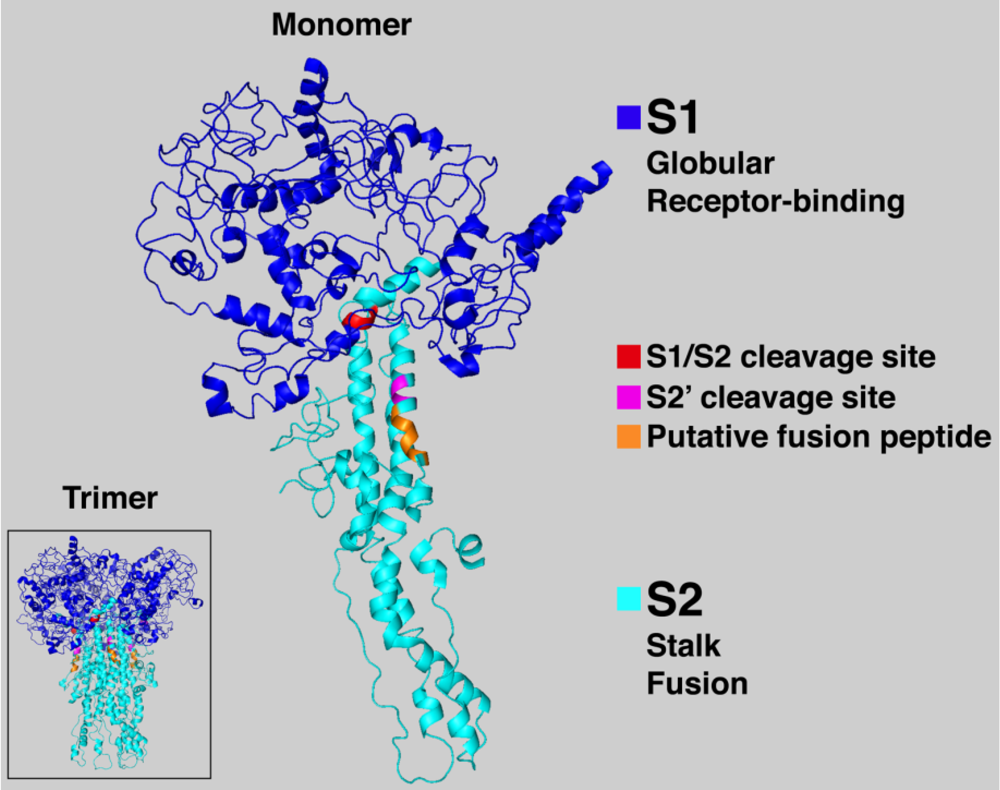
These models were subjected to molecular dynamics (MD) simulation to determine the extent to which glycan microheterogeneity impacts the antigenicity of the S glycoprotein. 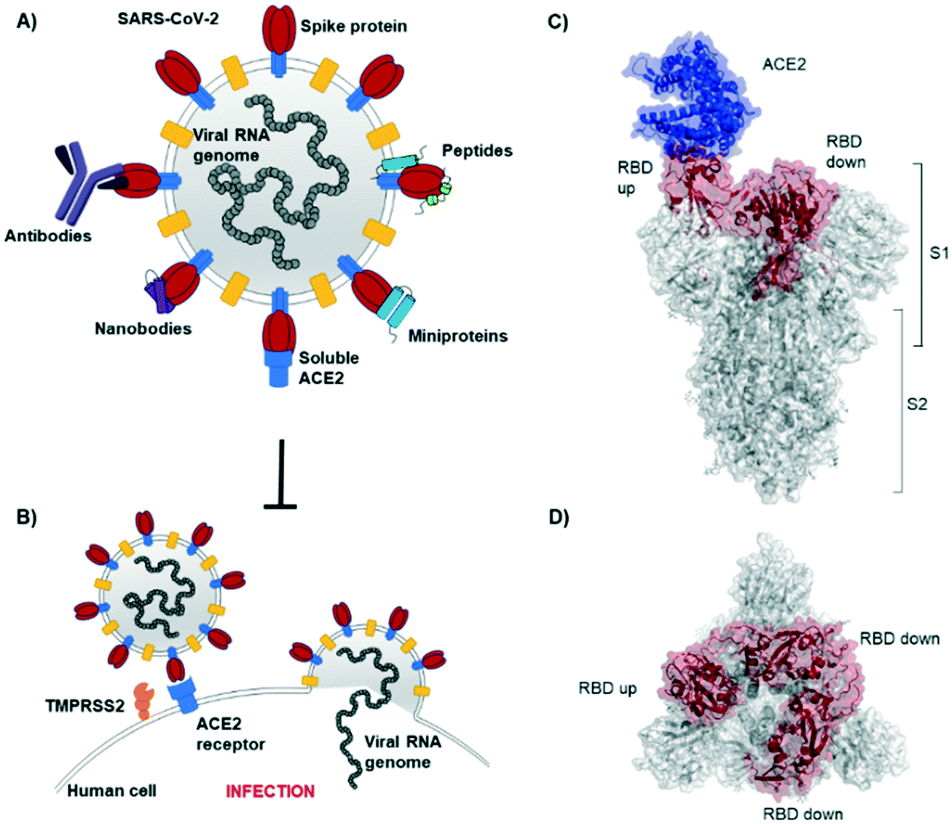
We also analyze structures for glycoforms representing those present in the nascent glycoproteins (prior to enzymatic modifications in the Golgi), as well as those that are commonly observed on antigens present in other viruses. Here we have generated 3D structures of glycoforms of the spike (S) glycoprotein from SARS-CoV-2, based on reported 3D structures and glycomics data for the protein produced in HEK293 cells.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
